How Accurate is the COVID-19 PCR Test Really?
A deep dive into Christian Drosten's COVID-19 PCR Primers...
Summary
Eight of ten SARS-CoV-2 PCR primers (Corman-Drosten et al 2020) are found in the SARS-CoV-2 genome.
All primers, by themselves, show at least a two third match to the human genome.
The primer “N_Sarbeco_R” shows a match of 19/20 base pairs to the human genome.
The real world impact of these results remain unclear, but many open questions arise!
Overview of my Seven Part Genomics Series:
Background
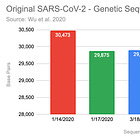
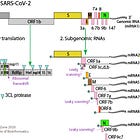
Wu et al. 2020 published three versions of the SARS-CoV-2 genome sequence. It is assumed, that based on these sequences, Christian Drosten designed the first PCR protocol for COVID-19. His infamous paper, made it through the peer review in less than 24h!
Noteworthy, ‘Drosten et al’ also did not have any live virus available for the calibration of his PCR essay, as they wrote: “In the present case of 2019-nCoV, virus isolates or samples from infected patients have so far not become available to the international public health community.”
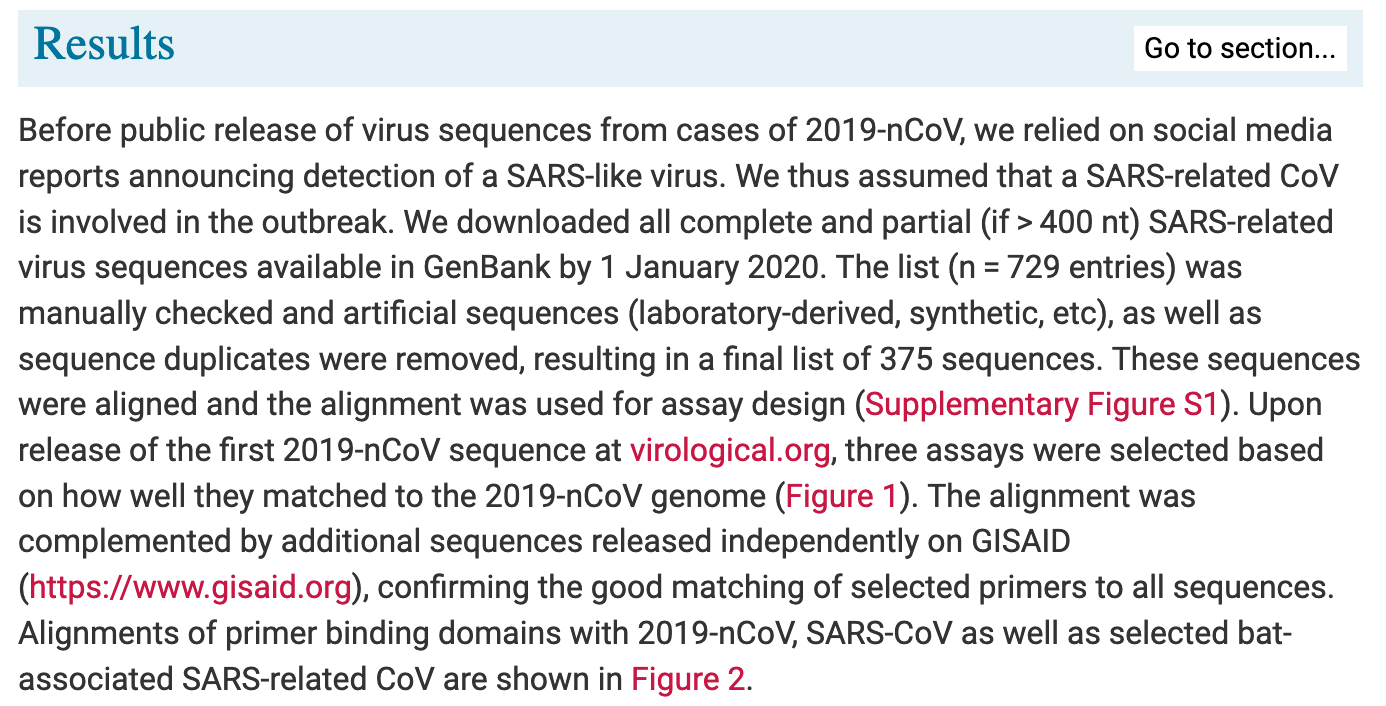
In the paper, he also lines out how exactly they designed and chose the primers:
So Drosten designed the PCR primers based on the assumption, that a SARS-related Coronavirus was involved in the outbreak in Wuhan, China - how could he know that? Well, Drosten, also identified the first version of SARS, in 2003, hence it suggests itself, that he would be on the lookout for a new version of the SARS virus which ‘mysteriously’ disappeared in 2004!
Primers
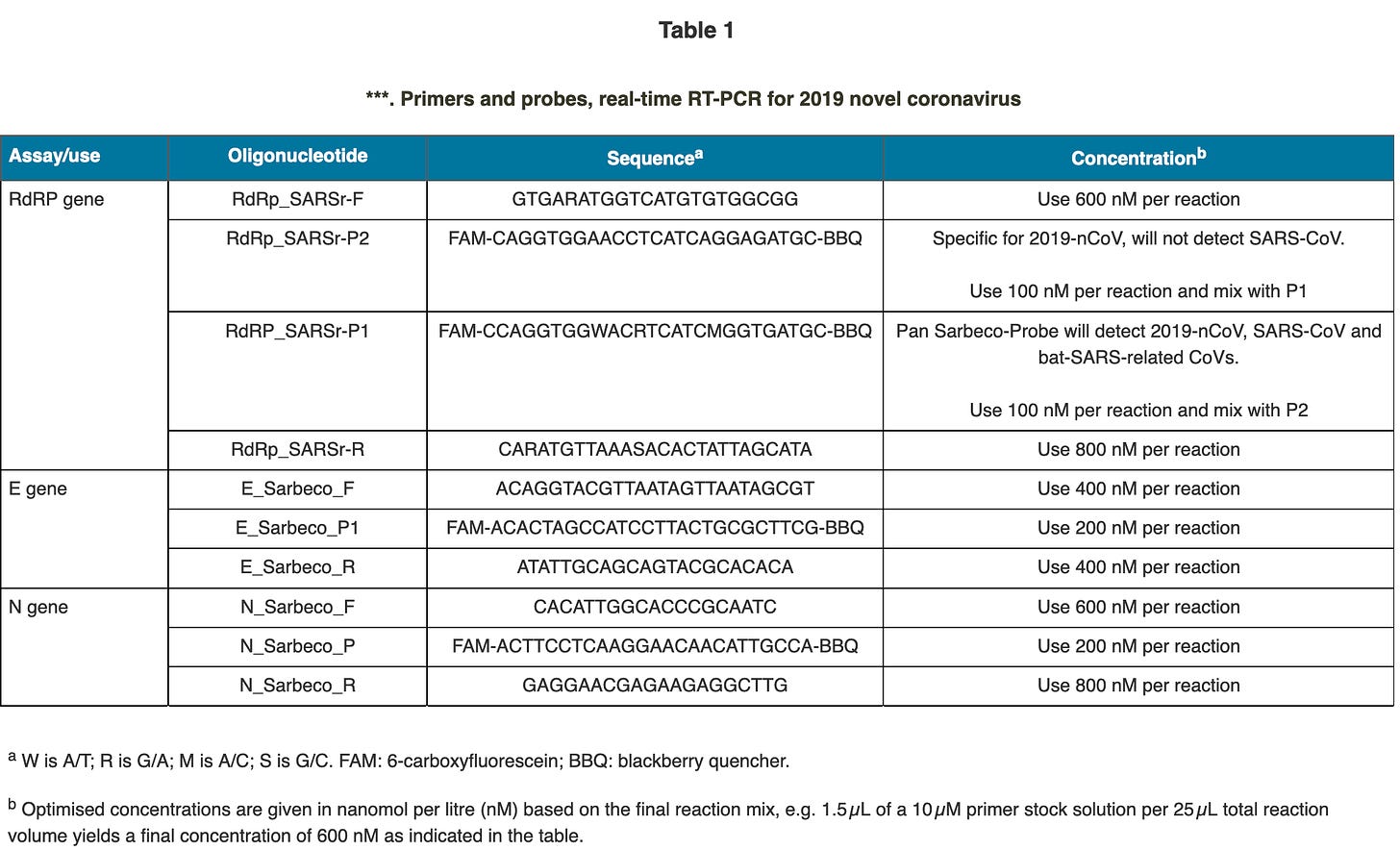
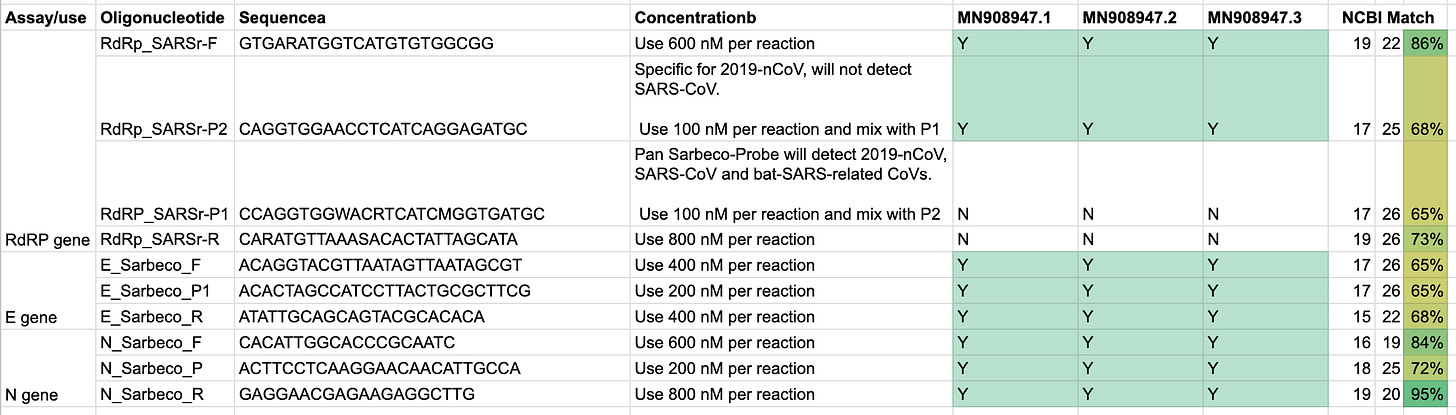
Drosten’s paper lists the final primers, that he suggests being used for detection of the allegedly “novel” coronavirus:
One of the questions that arose during my work on the genome sequencing of Wu et al’s SARS-CoV-2 genome was, if these primer sequences are actually present in the genome.
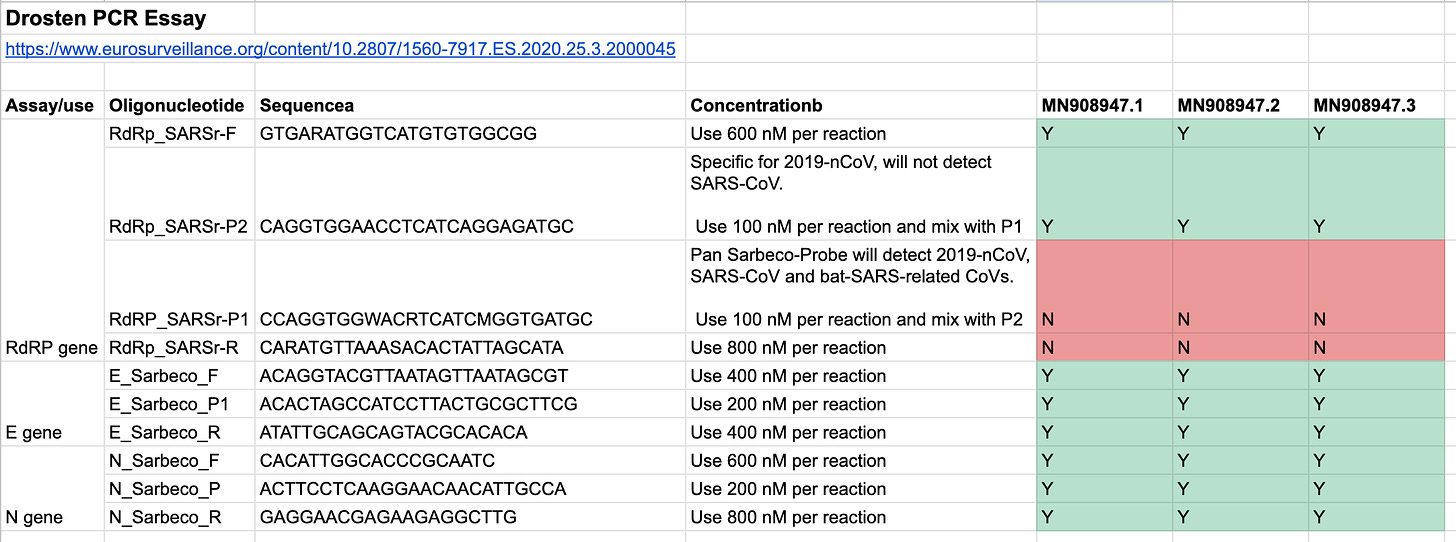
The short answer is yes, except two, let’s have a look:
All, except the RdRP_SARSr-P1 & RdRp_SARSr-R sequences can be found in the sequence. I have validated the presence via the commands documented in this script.
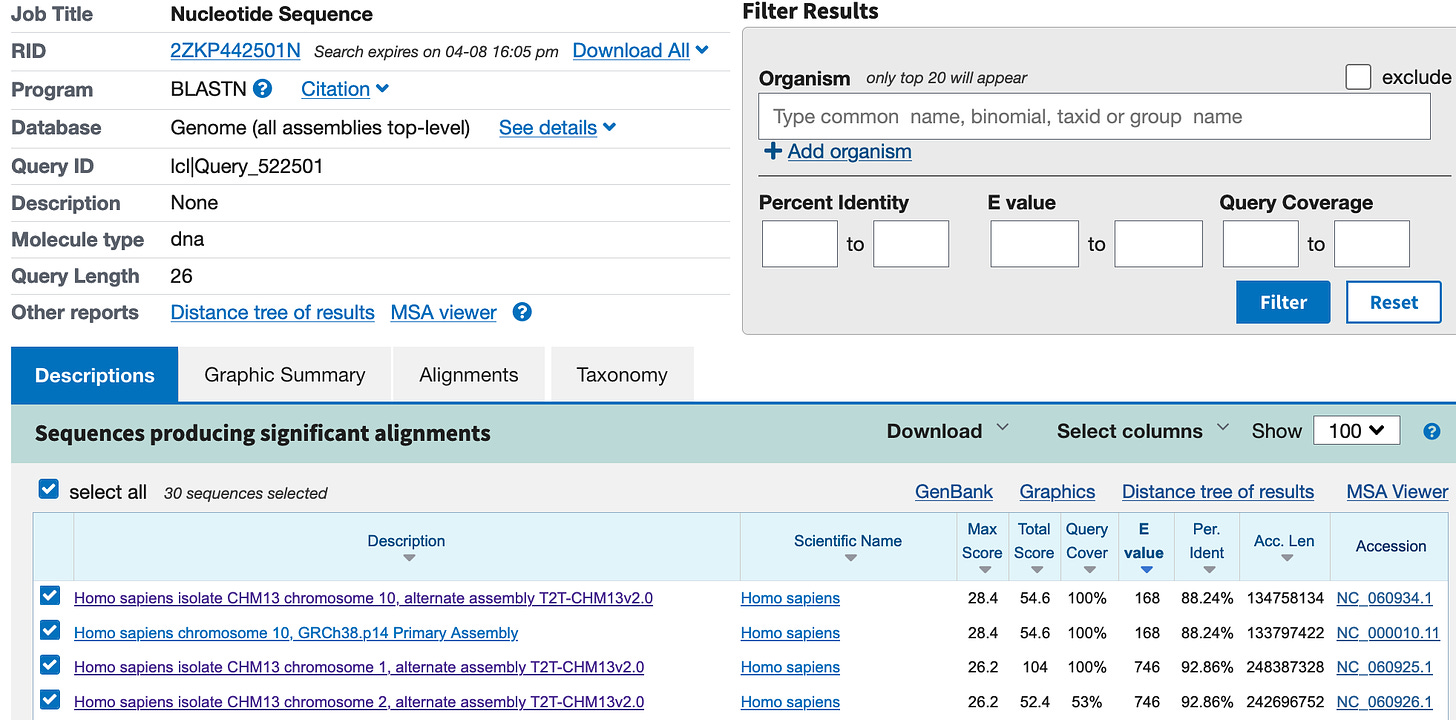
It’s unclear to me why two primers do not match, and what they are actually detecting. A quick blast search, via the online tool at NCBI, lists partial matches with the human genome for 17 of the 26 base pairs, but fails to match SARS1 or SARS2.
So this is interesting, the primers already show a high match to the human genome?!
Blasting All Ten Primers
So next, I ‘blasted’ all the primers with the above mentioned script. Here’s the result:
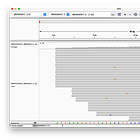
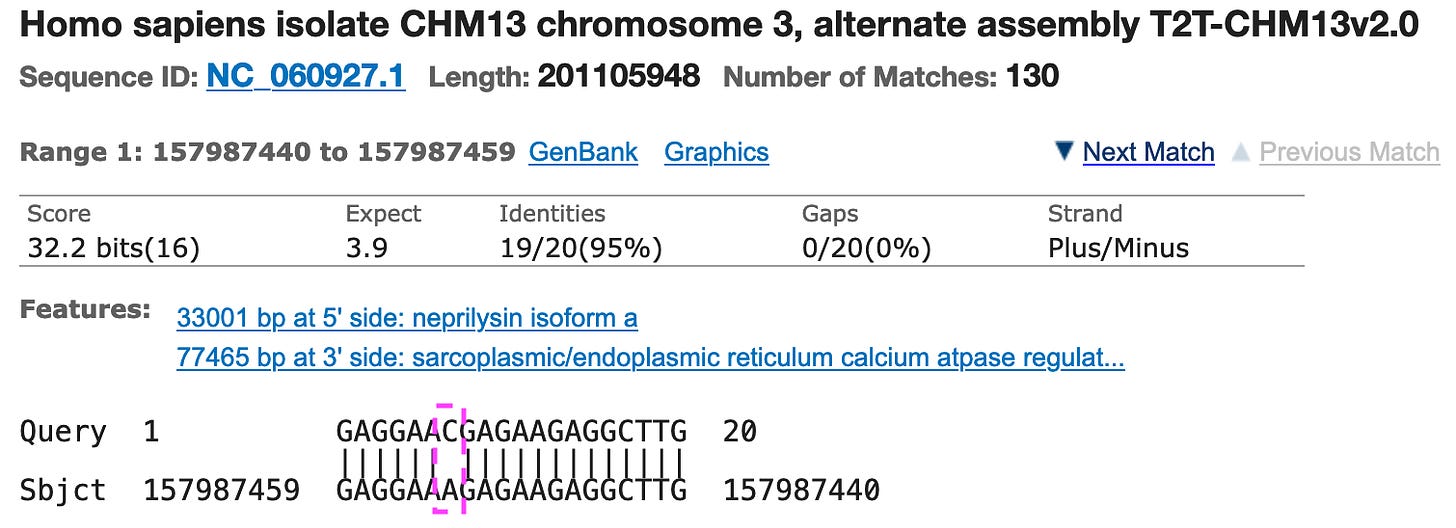
Most of the primers matched large parts of the human genome. The last primer “N_Sarbeco_R” matched the human genome for 19 of 20 base pairs!!
As you can see here, the full primer matches, except the C at position 7, here the human genome shows an A.
The other primers all show a relatively high match rate as well of over 60%+ to the human genome.
Here’s an example in which I show that for the “E_Sarbeco_R” primer, not only do we get a pretty high match of 82% to the human genome (with some translation errors in between), but once you align two strands of human DNA to the primer, it becomes actually a 100% perfect match. Now, I do not know if this case is possible in the PCR, and partial matches like this are possible or somehow guarded against.
Discussion
As shown above, not all PCR primers suggested by Drosten match the SARS-Cov-2 genome. It remains unclear why.
Partial matches, and in one case almost complete match to the human genome are found. This introduces the question, of how likely, simple errors in the PCR amplification process, could lead to positive results.
As far as I know, only a few hundred human genomes have actually been sequenced entirely. I would think that these genomes are not all the same, and what about the other 8 billion human genomes on this planet? How likely is it, that these primers match some of our human genomes? And what happens when the DNA is “denatured”, i.e. via all kind of influences, such as drugs, illnesses, bacteria, viruses, etc.?
To me, this shows that these COVID-19 PCR tests have not been validated properly (and clinically as well) and should not be used a determining factor for diagnoses or public health measures. At best, maybe as a guide for medical professionals to guide treatment of ill patients.
I did talk to some PCR experts, which assured me, that these primers would normally not turn the test positive at random - in ideal situations - and unless they are not run for too many cycles!
But still the close resemblance to the human genome sequences, makes me stay suspicious, especially if pieces of human genome could potentially turn the test positive!?
Let me know what you think in the comments!
Please like & share!
Update on the Mismatches
CARATGTTAAASACACTATTAGCATA
CAAATGTTAAASACACTATTAGCATA
CAAATGTTAAAAACACTATTAGCATA
W is A/T; R is G/A; M is A/C; S is G/C.
CCAGGTGGWACRTCATCMGGTGATGC
CCAGGTGGAACrTCATCAGGtGATGC
CCAGGTGGAACCTCATCAGGAGATGC
RdRP_SARSr-P1
pos 12, S instead of R
pos 21, T instead of A








I don't know about the US. But in Germany the ideal situation was never present. The labs did run double the amount of cycles that could give you a real idea if the person has more then a scrap of whatever the test looks for in his mouth or nostril. It cannot say if it is viable or dead. It can just say it's present. But with a too high number of cycles, we are talking trace amount. I know people with two positive tests and two negative ones on the same day. I don't know if the human genome match could potentially make false positives. But overtesting and a too high number of cycles sure can. Up to well over 90% false positives. Thats why monitoring healthy school children or otherwise symptomless people with PCR isn't a meaningful thing to do. My poor grandniece is becoming an IC nurse. The kid went six times into PCR quarantine, plus two times Corona infection, which not always led to a positive test right when it became interesting in terms of potential transmission. If this even was the goal.
To create numbers and fear, the test was fabulous. And fear is, what Dr. Drosten knows and likes best, he has done so with swine flu before. So yes I believe he made the test to create cases not to detect actual illness, but I cannot say, if the human genome matches can add to that.
Well of course they needed a test that would give positives to something that's never existed. Otherwise there would have been no plandemic.